DNA Extraction:
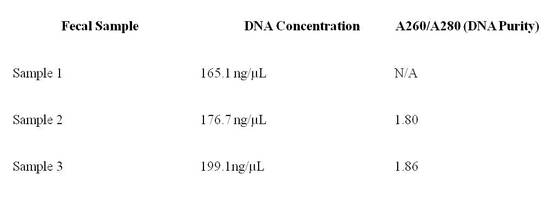
Table 1: Concentrations and purity of extracted DNA shown in table
above. These measurements were determined using a nanodrop spectrophotometer.
First PCR Reaction:
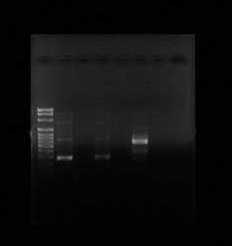
Only two samples of the PCR reaction were successfully amplified. Samples 2 and 4 both showed solid bands. Sample 2 is found in lane 1 and lane 2 and it appears to have approximately 300 base pairs. Sample 2 contained the 515F/806R gene fragments. Sample 4 is located in lane 5, and based on the amount the DNA travelled across the gel, it appears to approximately 550 base pairs long. Sample 4 contained the NS1/NS2 gene fragments. However, sample 6 never appeared on the gel.
Only two samples of the PCR reaction were successfully amplified. Samples 2 and 4 both showed solid bands. Sample 2 is found in lane 1 and lane 2 and it appears to have approximately 300 base pairs. Sample 2 contained the 515F/806R gene fragments. Sample 4 is located in lane 5, and based on the amount the DNA travelled across the gel, it appears to approximately 550 base pairs long. Sample 4 contained the NS1/NS2 gene fragments. However, sample 6 never appeared on the gel.
Figure 1: This gel shows results of first PCR reaction. Lane 2 and 4 both contained 515/806 gene fragments. Lane 6 contained the amplified NS1/NS2 gene fragments
Second PCR Reaction:
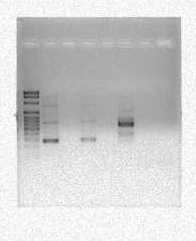
Lane 3 of the gel contained the 515F/806R gene fragments while lanes 6 and 7 contained the 5313F/1492R gene fragments. The sample in lane three appears to be approximately 300 base pairs long when compared to the DNA ladder in lane 1. Also, the samples in both lane 6 and 7 appear to be about 900 base pairs long when compared to the DNA ladder in lane 1. Lanes 4 and 5 do show any bands of DNA because the NS1/NS2 samples ran out so they were not present in this well.
Lane 3 of the gel contained the 515F/806R gene fragments while lanes 6 and 7 contained the 5313F/1492R gene fragments. The sample in lane three appears to be approximately 300 base pairs long when compared to the DNA ladder in lane 1. Also, the samples in both lane 6 and 7 appear to be about 900 base pairs long when compared to the DNA ladder in lane 1. Lanes 4 and 5 do show any bands of DNA because the NS1/NS2 samples ran out so they were not present in this well.
Figure 2: Results of second PCR which had annealing temperature of 60ᵒC instead of 55ᵒC. The first lane 2 contained 515/806 gene fragments. Lane 5 and 6 contained the 533/1492 gene fragments.
Third PCR Reaction:
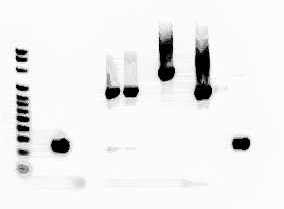
The band in Lane four contains NS1/NS2 gene fragment samples and is approximately 550 base pairs long as expected when compared to the DNA Ladder. The 533F13/1492R gene fragment samples were also run through this gel and the bright band seen in lane 6 of figure 3 is approximately 900 base pairs long.
The band in Lane four contains NS1/NS2 gene fragment samples and is approximately 550 base pairs long as expected when compared to the DNA Ladder. The 533F13/1492R gene fragment samples were also run through this gel and the bright band seen in lane 6 of figure 3 is approximately 900 base pairs long.
Figure 3: This gel was meant to make sure that the NS1/NS2 samples were successfully amplified in PCR. Lane 4 contains the NS1/NS2 gene fragments and Lane 7 contains the 533F/1492R gene fragments.
Transformation Colony Counts:
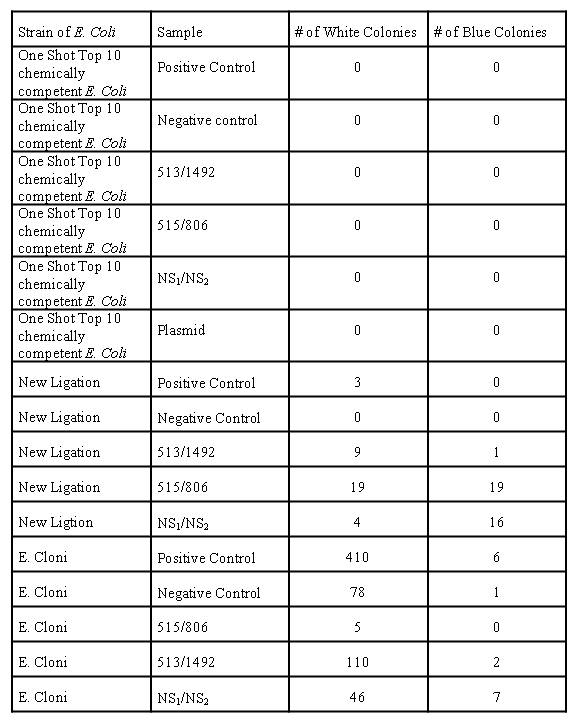
Table 2: Number of white and blue colonies present on each plate
Table 2 shows that no colonies grew on plates containing the One Shot TOP10 chemically competent E. coli. For the Lucigen E. cloni bacteria, colonies grew on every plate including the negative control. The positive control plate that had Lucigen E. cloni cells had the greatest number of white and blue colonies. The plates that contained the NEB 5-alpha competent E. coli and the higher quality ligation reaction had the best results. No colonies grew on the negative control plate that contained cells with the higher quality ligation; however the number of blue colonies for the plates containing the NEB 5-alpha competent E. coli cells with the higher quality ligation was much higher compared to the other plates.
Sequencing and BLAST:
MC1 (515F/806R): Neisseria bacilliformis
MC3 (515F/806R): Neisseria flava
MC5 (1492/533533F/1492R): Neisseria flava Uncultured organism clone MC042313 small subunit ribosomal RNA gene, partial sequence
MC7 (NS1NS2): Mucor racemosus
MC1 (515F/806R): Neisseria bacilliformis
MC3 (515F/806R): Neisseria flava
MC5 (1492/533533F/1492R): Neisseria flava Uncultured organism clone MC042313 small subunit ribosomal RNA gene, partial sequence
MC7 (NS1NS2): Mucor racemosus